7. Metabolic Pathways
Many excellent reviews are available covering metabolic pathways in methanotrophs, e.g. (Anthony, 1982; Chistoserdova, 2011; Chistoserdova and Lidstrom, 2013; Dalton, 2005; Hanson and Hanson, 1996; Trotsenko and Murrell, 2008). This section will briefly describe known pathways and provide links for further reading. Please refer to Fig. 1 (overview, from the Introductory section). A set of slides on methylotrophic bacteria in general are available on Prof. Chris Anthony’s website that are useful as a succinct summary as well as a historical guide.
7.1 Methanol, formaldehyde, and formate oxidation
Methanol is generally oxidized to formaldehyde by the classical periplasmic MxaF-type methanol dehydrogenase, an α2β2 tetramer containing the cofactor PQQ and calcium ions. Biochemical information on this enzyme is mostly from non-methanotrophic methylotrophs (Anthony, 2004), although purification has been reported from several methanotrophic strains (Grosse et al., 1998; Grosse et al., 1997; Parker et al., 1987; Patel et al., 1972; Patel and Felix, 1976; Patel et al., 1978). Immunogold labeling has localized this enzyme to the internal membranes in a gamma-proteobacterial methanotroph, a Methylomonas strain (Fassel et al., 1992). The verrucomicrobial methanotrophs do not have genes for the MxaF-type methanol dehydrogenase, but instead have genes for the related PQQ-linked enzyme, XoxF (Khadem et al., 2012). This enzyme appears to require lanthanides in place of calcium and it has been reported that the verrucomicrobial methanotrophs require lanthanides for growth (Pol et al., 2013).
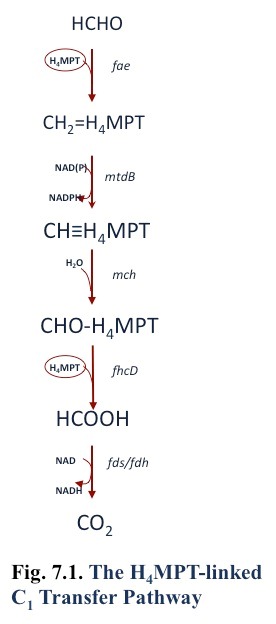
Formaldehyde may be oxidized to formate by the tetrahydromethanopterin pathway [Fig. 7.1; (Chistoserdova et al., 1998)], or in the gamma-proteobacterial methanotrophs by the cyclic ribulose monophosphate pathway (Anthony, 1982; Vorholt, 2002)(Anthony 1982), or both. It is possible that cytochrome- or NAD-linked formaldehyde dehydrogenases are also involved in formaldehyde oxidation (Vorholt, 2002; Zahn et al., 2001). However, the genes encoding a membrane-associated quinoprotein formaldehyde dehydrogenase in Methylococcus capsulatus Bath (Zahn et al., 2001) are not found in the genomes of other methanotrophs. In addition, one report of NAD-linked formaldehyde dehydrogenase in Methylococcus capsulatus Bath was shown to be due to combined activity of methanol dehydrogenase (which has formaldehyde dehydrogenase activity) and methylene tetrahydromethanopterin dehydrogenase (Adeosun et al., 2004).
Formate is oxidized to CO2 by formate dehydrogenase, and it is common for methanotroph genomes to contain genes for multiple types, including NAD-linked and dye-linked enzymes. Purification has been reported from Methylosinus trichosporium OB3b of an NAD-linked enzyme (Jollie and Lipscomb, 1991; Yoch et al., 1990) and a ferredoxin-linked enzyme (Chen and Yoch, 1989).
7.2 Serine Cycle
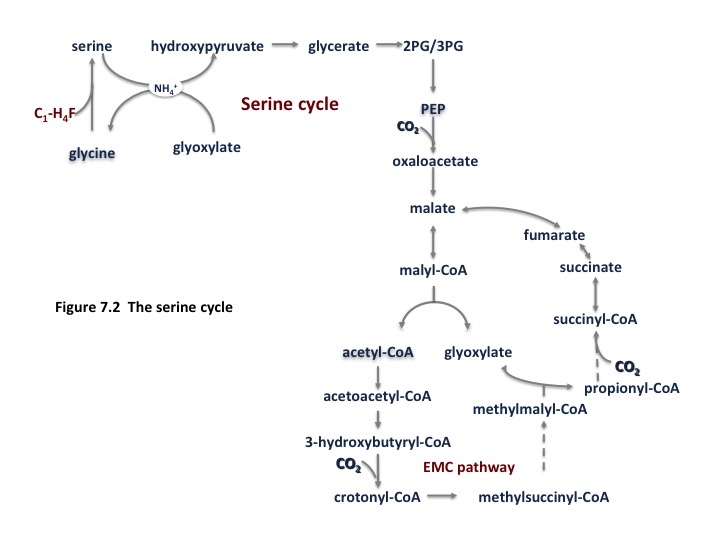
The alpha-proteobacterial methanotrophs utilize the serine cycle for formaldehyde assimilation. An abbreviated form of the cycle is shown in Fig. 7.2 and a complete version can be found in (Anthony, 2011). In this cycle, methane is converted to formate by the reactions noted above, which is converted to methylene tetrahydrofolate (H4F) via a standard H4F pathway. Methylene H4F condenses with glycine to generate serine. The remainder of the cycle is involved in regeneration of the glycine acceptor. The equation for generating one phosphoglycerate molecule is:
1.5 CO2 + 1.5 HCHO + 1.5 NADH + 1.5 ATP + 0.67 FAD ==>3-phosphoglycerate + 1.5 NAD + 1.5 ADP + 0.5 Pi +0.67 FADH
7.3 Ribulose Monophosphate Cycle
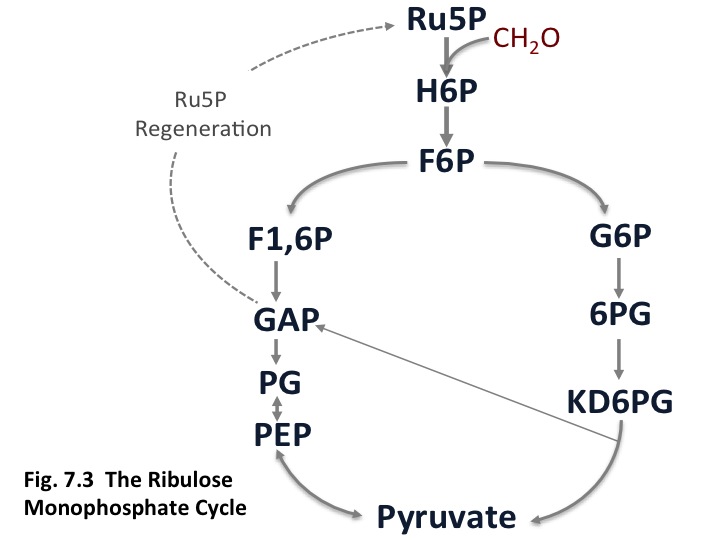
The gamma-proteobacterial methanotrophs use the ribulose monophosphate cycle. In this pathway [An abbreviated form of the cycle is shown in Fig. 7.3 and a complete version can be found in (Anthony, 1982)]. In this case, formaldehyde is condensed with ribulose monophosphate to create a hexulose phosphate, which is converted to fructose-6-phosphate. Two routes are possible to convert fructose-6-phosphate to pyruvate, the Entner-Douderoff (EDD) pathway, shown on the right, and the EMP pathway (glycolysis), shown on the left. Although enzyme activities suggested that the EDD pathway was the route that occurred in methanotrophs, it has recently been shown by 13C-labeling in a gamma-proteobacterial Methylomicrobium strain, that the dominant pathway for generating pyruvate is the EMP pathway (Kalyuzhnaya et al., In press). In the latter case, the reaction for generating a 3PGA is:
3 HCHO + NAD + 0.67 ADP + 5/3 Pi ==> PGA + NADH + 0.67 ATP
A comparison to the serine cycle shows that this pathway is much more efficient.
7.4 Calvin-Benson-Bassham Cycle (the Ribulose Bisphosphate Cycle)
The verrucomicrobial methanotrophs utilize the classical Calvin-Benson-Bassham cycle , oxidizing methane to CO2 for energy and incorporating CO2 into cell material by this cycle (Khadem et al., 2011).
Some methanotrophs, such as Methylococcus capsulatus Bath contain genes for all three assimilatory pathways, but evidence to date suggests that the RuMP pathway dominates carbon assimilation (Stanley and Dalton, 1982).
7.5. Respiration
|
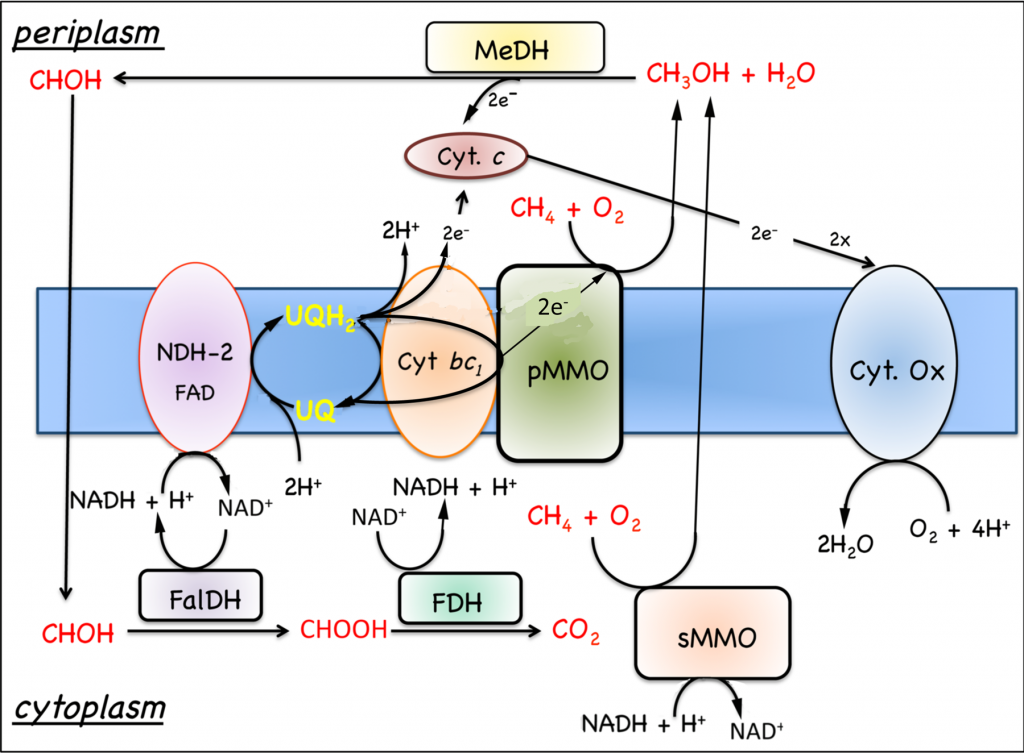
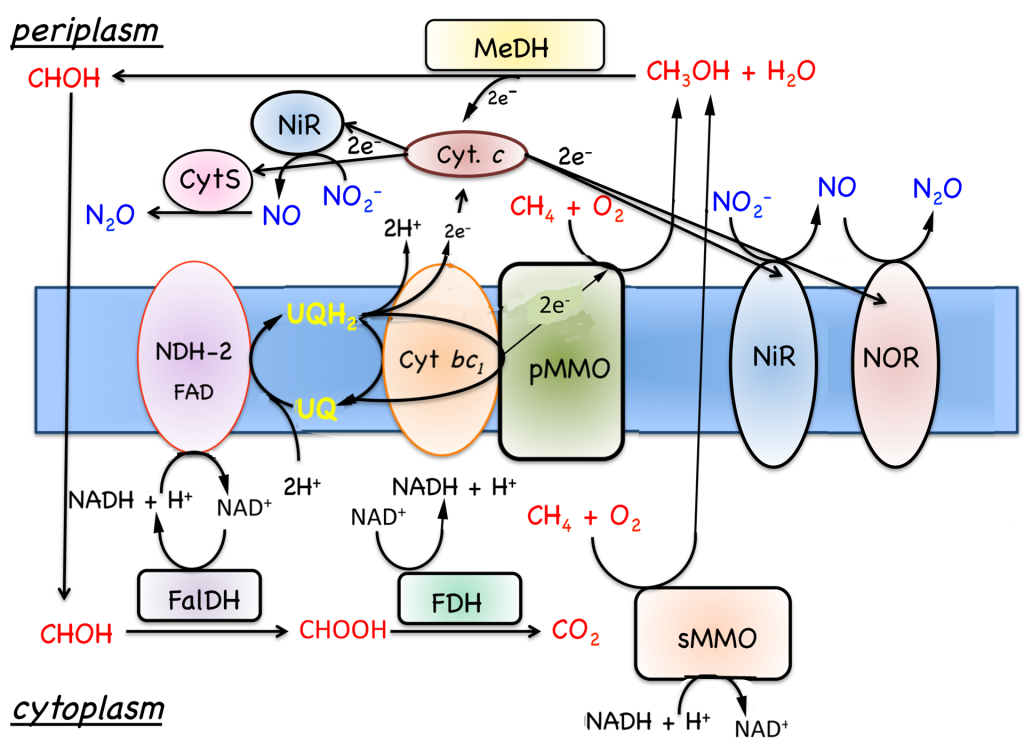
Figure. 7.4. Model for the oxidation of methane to carbon dioxide in M. capsulatus Bath. Substrate oxidation enzymes are represented by rectangles and respiratory components by ovals. Abbreviations: cyt., cytochrome; FalDH, formadehyde dehydrogenase; FDH, formate dehydrogenase; MeDH, methanol dehydrogenase; Ox, oxidase; pMMO, membrane-associated methane monooxygenase; UQ, ubiquinols; sMMO, soluble methane monooxygenase. |
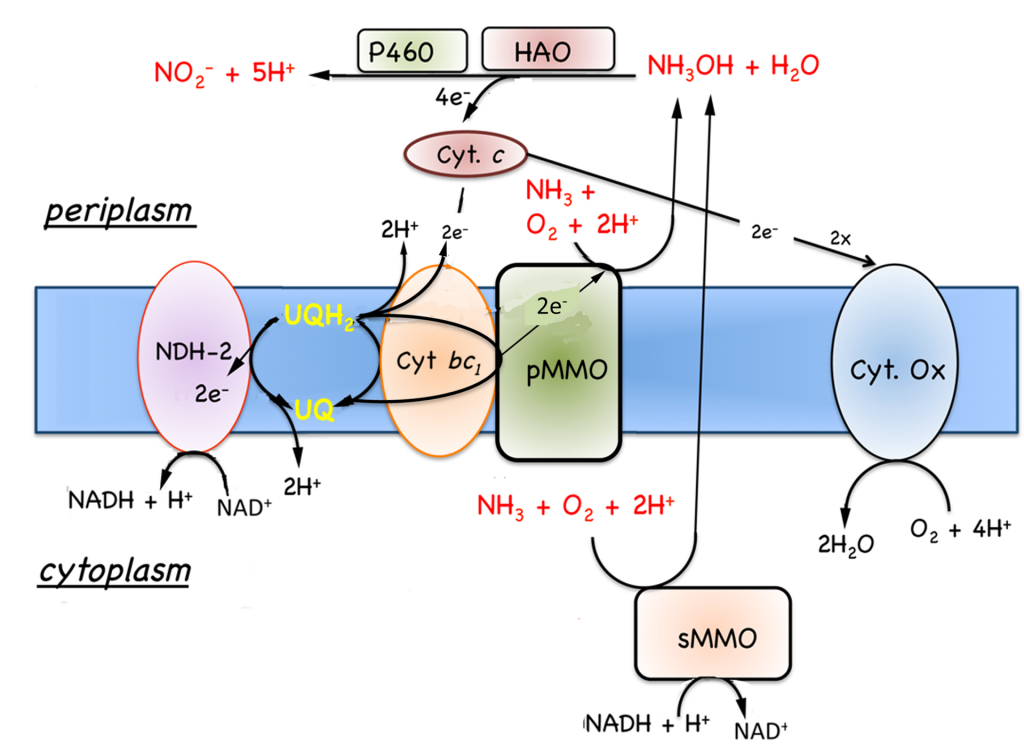
7.5.1 Oxidation of growth substrates, methane and methanol. As noted in Section 7.1, obligate aerobic methanotrophs oxidize methane or methanol to CO2 via a series of two electron steps with methanol, formaldehyde, and formate as intermediates (Fig. 7.4). The following discussion will focus on respiration in Methylococcus capsulatus Bath, since this is the best-characterized system (DiSpirito et al., 2004; Ward et al., 2004) and genomic analysis predicts similar respiratory chains in many other aerobic methanotrophs. As discussed in Sections 6.2 and 6.2, the oxidation of methane to methanol is an energy-dependent reaction and the electron donor for methane oxidation is thought to be either NADH + H+, in cells expressing the soluble methane monooxygenase (sMMO)(Colby and Dalton, 1978, 1979), or ubiquinol in cells expressing the pMMO (DiSpirito et al., 2004). The oxidation of methanol by the methanol dehydrogenase is coupled to a periplasmic mono-c-type cytochrome, cytochrome cL (Ambler et al., 1986; Anthony, 1982; Ward et al., 2004) and the oxidation of formate and formaldehyde by formate and formaldehyde dehydrogenases or other enzyme systems described in Section 7.1 generates NAD(P)H (Anthony, 1982; Attwood and Quayle, 1984; Jollie and Lipscomb, 1991; Vorholt, 2002; Zahn et al., 2001). Based on the growth
substrates one would predict a very limited or simple respiratory chain consisting of a complex 1 or NAD+-dehydrogenase, complex 3 or a cytochrome bc1-type complex and complex IV or dioxygen dependent terminal electron acceptor, all of which have been identified in genome sequences. However, genomic and biochemical studies have identified a surprising number of respiratory components in aerobic methanotrophs, which go beyond the metabolic capabilities predicted from the growth substrates of this microbial group (Boden et al., 2011; Chen, 2010; Dam et al., 2012; DiSpirito et al., 2004; Khmelenina et al., 2013; Kits et al., 2013; Stein et al., 2011; Ward et al., 2004).
7.5.2 Oxidation of ammonia and hydrogen. In addition to the known growth substrates, methane and methanol, methanotrophs will also oxidize molecular hydrogen and ammonia. Ammonia is oxidized to hydroxylamine in an energy-dependent reaction by both MMOs (Dalton, 1977), and hydroxylamine is oxidized by either an octyl-c-heme type cytochrome, hydroxylamine oxidoreductase (HAO) (Campbell et al., 2011) or via cytochrome P460 (Fig. 7.5) (Zahn et al., 1994). No putative HAO has been isolated from an aerobic methanotroph, so the electron acceptor for this enzyme is not known. In aerobic ammonia oxidizing bacteria the electron acceptor for HAO is a tetraheme cytochrome
(Andersson et al., 1986; Arciero et al., 1991; Hooper et al., 2004). However, no tetra-c-heme cytochromes have been identified in methanotrophs, so if these putative HAOs do oxidize hydroxylamine, they utilize a difference class of electron acceptors. Cytochrome P-460 utilizes a mono-heme c-type cytochrome, probably cytochrome cL or cytochrome c’, as an electron acceptor (Fig. 7.5) (Zahn et al., 1996; Zahn et al., 1994).
The hydrogenases identified in AMOB are NAD+-linked (Ward et al., 2004). Thus, the oxidation of both ammonia and hydrogen would potentially contribute to the proton gradient and supply metabolic energy, although neither substrate has been shown to serve as a sole energy source.
7.5.3 Oxygen-, nitrite- and nitrous oxide-dependent terminal electron acceptors.
Methane is oxidized to methanol via a monooxygenase reaction and dioxygen must be provided either externally or internally via reduction of nitrite (Ettwig et al., 2010; Lipscomb, 1994). Thus, oxygen is required for growth on methane. However, methanotrophs grow under a variety of aerobic, microaerophillic and even anaerobic conditions. The variety of terminal oxidases reflects this diversity and most methanotrophs contain a aa3-type oxidase as well as high affinity heme d-type oxidases (Dispirito et al., 1994; Ward et al., 2004).
In addition to oxygen-dependent most methanotrophs examined contain both Cu and cytochrome cd-type dissimilatory nitrite reductases and putative nitrous oxide reductases (Fig. 7.6); (Boden et al., 2011; Chen, 2010; Dam et al., 2012; DiSpirito et al., 2004; Khmelenina et al., 2013; Kits et al., 2013; Stein et al., 2011; Ward et al., 2004). M. capsulatus Bath has also been shown to contain 6 high molecular mass multi-c-heme cytochromes similar to high molecular mass cytochromes associated with metal reduction (Bergmann et al., 1999; Ward et al., 2004). However, the function of these high molecular mass cytochromes has not been determined and have only been detected in M. capsulatus Bath.
7.5.4 Conclusions. The function of a large number of respiratory components identified in the sequenced genomes of AMOB have yet to be identified. Some of the additional complexity is found in the oxidation of non-growth substrates, and the results suggest the metabolic capabilities of AMOB are greater than generally predicted from the known metabolism.
7.6. Nitrogen Metabolism
The ability of methanotrophs to fix dinitrogen (N2) was originally thought to be mainly confined to type II (alphaproteobacterial) and type X methanotrophs including Methylosinus trichosporium and Methylococcus capsulatus Bath, respectively. However, PCR assays for nifH, encoding the iron protein of nitrogenase which is highly conserved in all nitrogen fixing bacteria, and acetylene reduction assays, has revealed the potential for N2 fixing ability among some Type I (gammaproteobacterial) methanotrophs such as Methylomonas and Methylobacter. The structural genes for nitrogenase of Methylococcus capsulatus Bath (nifH, nifD, and nifK) were shown to be contiguous as they are in other N2 fixing bacteria. Genome analysis shows that this contiguous region extends to include other nif genes such as nifN and nifX, which are involved in synthesis of the nitrogenase iron-molybdenum cofactor (Auman et al., 2001; Murrell and Dalton, 1983b; Trotsenko and Murrell, 2008).
Type I (gammaproteobacterial) methanotrophs generally assimilate ammonium (NH4+) by reductive amination of alpha-ketoglutarate or pyruvate using the enzymes glutamate dehydrogenase or alanine dehydrogenase respectively. Type II (alphaproteobacterial) methanotrophs use the glutamate cycle and the enzymes glutamine synthetase (GS) and the glutamine-oxoglutarate amidotransferase (GOGAT, also known as glutamate synthase). In Type I methanotrophs grown on medium containing ammonia, the reductive amination of pyruvate (via alanine dehydrogenase (in M. capsulatus Bath) or alpha-ketoglutarate (via glutamate dehydrogenase) was prevalent under high ammonia growth conditions. In contrast, when grown under ammonium limitation (<0.5 mM) or on medium containing nitrate (in the absence of ammonium) these methanotrophs assimilated ammonia via the glutamate cycle. Four predicted ammonium transporters have been identified in the genome of M. capsulatus Bath (Murrell and Dalton, 1983a; Trotsenko and Murrell, 2008).
The GS purified from M. capsulatus Bath is a homododecameric enzyme (617 kDa) with a subunit molecular mass of 50 kDa. In M. capsulatus Bath, glutamine synthetase is subject to regulation by the adenylylation/deadenylylation cascade, exhibiting a rapid switch off of GS activity when excess ammonia is added to nitrogen-limited cultures. At concentrations of >0.5 mM NH4+ in the medium, GS exists in the non-active adenylylated form. Expression of the structural gene for GS from M. capsulatus Bath, glnA, is regulated via the Ntr system found in enteric bacteria. The levels of GS/GOGAT, glutamate, and alanine dehydrogenases in methanotrophs are also dependent on the nitrogen content in growth medium. The GOGAT of type I methanotrophs requires NADPH and is rather unstable, whereas in type II methanotrophs this enzyme is more stable and specific for the cofactor NADH (Trotsenko and Murrell, 2008).